About Article
Molecular Insights into Arylsulfatase B Mutation-Induced Instability in Mucopolysaccharidosis Type VI
Volume 1, No. 1 · 2026
Highlights
- The study functionally annotated 92 hypothetical proteins from Listeria monocytogenes serotype 4b using an integrative in silico workflow.
- Sequence-based analyses used BLAST, Pfam, InterPro, motif analysis, and multiple sequence alignment to classify proteins into major functional groups.
- Thirteen selected hypothetical proteins were modeled structurally using SWISS-MODEL and analyzed by PyMOL, DALI-Lite, and ProFunc.
- Predicted functions covered hydrolases, transferases, transporters, kinases, stress-response proteins, membrane proteins, DNA-binding proteins, phosphoesterases, and ATP-binding proteins.
- The work improves genome annotation quality for L. monocytogenes and proposes experimentally testable functions for previously uncharacterized proteins.
Abstract
The article addresses the large number of hypothetical proteins that remain uncharacterized in bacterial genomes despite rapid advances in sequencing.
Using Listeria monocytogenes serotype 4b as a model pathogen, the study retrieved 92 hypothetical proteins and analyzed them through sequence similarity search, conserved-domain identification, motif analysis, and multiple sequence alignment.
Functional classification was primarily supported by BLAST, Pfam, and InterPro analyses, while selected proteins underwent homology modeling with SWISS-MODEL followed by structural validation and comparative analysis with PyMOL and DALI-Lite.
Structure-based functional inference was further supported by conserved-residue mapping and ProFunc analysis, which highlighted catalytic residues, ligand-binding regions, metal-binding sites, and other conserved functional motifs.
Overall, the study proposes roles for many previously uncharacterized proteins in enzymatic activity, transport, transcriptional regulation, stress adaptation, nucleotide metabolism, and membrane biology, and identifies candidates for future experimental validation.
Keywords
Article Overview
This article focuses on hypothetical proteins in Listeria monocytogenes, emphasizing that genomic sequencing has outpaced experimental functional annotation and left many predicted proteins uncharacterized.
The study frames functional annotation as important both for improving genome quality and for identifying potential biomarkers, therapeutic targets, and previously unknown biological pathways.
Because Listeria monocytogenes serotype 4b is a major food-borne pathogen with high mortality and contains 92 hypothetical proteins in its genome, the paper applies sequence-based and structure-based bioinformatics approaches to assign plausible functions to these proteins.
1. About the Article
This is a research article in Clinical & Molecular Biomedicine centered on genome-wide annotation and structural modeling of hypothetical proteins in Listeria monocytogenes.
The paper is authored by Shama Khan of the South African Medical Research Council / University of the Witwatersrand, with contact emails and an ORCID listed on the first page.
- Journal: Clinical & Molecular Biomedicine
- Volume/Issue: Vol. 1, No. 1
- Header year: 2026
- Title: Genome-wide annotation and structural modeling of hypothetical proteins in Listeria monocytogenes
- Author: Shama Khan
- Affiliation: South African Medical Research Council, Vaccine and Infectious Diseases Analytics Research Unit (VIDA), Faculty of Health Sciences, University of the Witwatersrand, Johannesburg, South Africa
- Corresponding author: Shama Khan
- Corresponding emails: Shama.Khan@wits-vida.org; Shama.Khan@wits.ac.za
- ORCID: 0000-0002-0874-5029
- License: Creative Commons CC-BY 4.0
- Open-access status: Open Access
- Header page range on first page: pp. 1–25
- Visible article page range in uploaded PDF: 1–12
2. Introduction
The introduction explains that hypothetical proteins are predicted from nucleotide sequence but lack direct experimental protein-level validation, and that many sequenced genomes still contain large fractions of such uncharacterized open reading frames.
The article argues that assigning functions to hypothetical proteins is essential for improving genome annotation, discovering new folds and domains, identifying biomarkers and therapeutic targets, and expanding understanding of biological pathways.
For Listeria monocytogenes, the paper emphasizes its clinical importance as a food-borne pathogen with high mortality and notes that serotype 4b contains 92 hypothetical proteins whose functions are poorly characterized.
- Organism of interest: Listeria monocytogenes serotype 4b
- Genome size noted: approximately 3 Mb
- Total nucleotides reported: 2,912,690
- Total protein genes reported: 2,766
- Total RNA genes reported: 85
- Hypothetical proteins highlighted in the genome: 92
3. Materials and Methods
The workflow begins with retrieval of 92 hypothetical protein sequences from the PIR website, saving them in FASTA format and tabulating accession numbers, molecular mass, length, and proposed functions.
Function prediction relied on sequence similarity searches with BLASTP, multiple sequence alignment with ClustalW, conserved domain and motif annotation with Pfam and InterPro, and structural-function prediction using SWISS-MODEL, PyMOL, DALI-Lite, and ProFunc.
Structural analysis included homology modeling of selected proteins, superposition and RMSD analysis in PyMOL, and comparison of modeled proteins with known templates to identify conserved active-site and ligand-binding residues.
- Data source for hypothetical proteins: PIR website
- Number of hypothetical proteins downloaded: 92
- Sequence format used for downstream analysis: FASTA
- Sequence tools: BLASTP; ClustalW; Pfam; InterPro
- Structure modeling server: SWISS-MODEL
- Visualization and superposition tool: PyMOL
- Structure-comparison tool used in the study: DALI-Lite
- Structure-based function inference tool: ProFunc
2.1. Data mining
The authors analyzed the Listeria monocytogenes genome at the PIR website and downloaded 92 hypothetical protein sequences.
Protein sequences were recorded with accession number, length, molecular mass, and proposed function, and saved in FASTA format for further analysis.
2.2. Sequence similarity search
BLASTP against public protein databases was used to identify related proteins and infer possible function from homologs of known activity.
Multiple sequence alignment with ClustalW was used to analyze conserved residues and support function prediction.
2.3. Structure Prediction
Homology modeling was used to infer three-dimensional structures of selected hypothetical proteins through template identification, alignment, fragment assembly, refinement, and energy minimization.
SWISS-MODEL was used to generate models, and 13 hypothetical proteins were selected for detailed structural analysis.
2.4. Function prediction
The article discusses several complementary function-prediction strategies, including clustering of gene-expression profiles, protein-protein interaction networks, motif/signature databases, and structure-based inference.
In practice, this study specifically used Pfam, InterPro, and ProFunc for functional interpretation.
2.5. Visualization
PyMOL was used for superposition of modeled proteins with templates, secondary-structure-based alignment, RMSD calculations, and residue-level visualization.
2.6. Structure analysis
The authors describe structure-based comparison methods for identifying remote homology and conserved functional sites, and select DALI-Lite for structural comparison in this work.
4. Results and Discussion
Sequence-based analysis classified all 92 hypothetical proteins into 13 broad categories, including hydrolase, transferase, transporter, RNA binding, hydrolase acting on carbon–nitrogen bonds, kinase, response to stress, integral membrane, DNA binding, phosphoesterase, ATP binding, others, and unknown.
Structural modeling was successfully carried out for 13 hypothetical proteins, and the models were compared with experimentally characterized templates to identify conserved catalytic residues, metal-binding sites, ligand-binding motifs, and likely molecular functions.
The study concludes that sequence-based classification plus structure-based analysis substantially improves annotation quality for Listeria monocytogenes, although several predictions with low sequence identity or high RMSD require cautious interpretation and future biochemical validation.
- Total hypothetical proteins analyzed: 92
- Number of proteins structurally modeled in detail: 13
- Major classes reported: hydrolase; transferase; transporter; RNA binding; C–N hydrolase; kinase; stress response; integral membrane; DNA binding; phosphoesterase; ATP binding; others; unknown
3.1. Sequence-based Function Prediction
The article grouped all 92 hypothetical proteins into major functional categories based primarily on sequence similarity, motif composition, and conserved-domain evidence.
Hydrolases and transferases were among the largest groups, while additional categories included transporters, RNA-binding proteins, kinases, stress proteins, membrane proteins, DNA-binding proteins, phosphoesterases, ATP-binding proteins, other singleton functions, and unknown proteins.
- Hydrolase examples: C1L0U3, C1L0R8, C1KZQ3, C1KZE1, C1KYZ2, C1KYY1, C1KY45, C1KY43, C1KXZ5
- Transferase examples: C1L2K8, C1L1Z7, C1L0P3, C1L0L0, C1L024, C1KYI4, C1KXY4, C1KVW1, C1KVA0
- Transporter examples: C1L300, C1L1R1, C1L164, C1L0K2, C1KZ81, C1KZ17, C1KYZ6, C1KY61, C1KY46
3.2. Structure-based Function Prediction
Templates were identified for thirteen proteins, and structural superimposition plus residue conservation analysis was used to infer active sites and likely biochemical roles.
Some predictions are strongly supported by high sequence identity and low RMSD, while others are marked as tentative due to weak structural agreement.
3.2.1. C1L1U8
Model built for residues 6–555 with 81.09% sequence identity to template and RMSD of 0.1 Å.
Conserved Zn2+-binding residues and catalytic residues support annotation as a nuclease, possibly with 5′–3′ exonuclease activity.
- Model size: residues 6–555
- Sequence identity: 81.09%
- RMSD: 0.1 Å
- Predicted function: nuclease / possible 5′–3′ exonuclease
3.2.2. C1KYM2
Model built for residues 2–259 with 37.31% sequence identity and RMSD of 1.359 Å.
Conserved catalytic triad residues and binding-cavity features support hydrolase activity acting on carbon–nitrogen bonds.
- Sequence identity: 37.31%
- RMSD: 1.359 Å
3.2.3. C1KYY1
Model built for residues 1–439 with 48.23% sequence identity and RMSD of 0.7 Å.
Conservation of the catalytic dyad and ligand-contact residues suggests a deoxynucleotide triphosphate triphosphohydrolase (dNTPase).
- Sequence identity: 48.23%
- RMSD: 0.7 Å
- Predicted function: dNTPase
3.2.4. C1L0M8
Model built for residues 5–105 with 40.78% sequence identity and RMSD of 0.5 Å.
Conserved DNA-binding residues suggest a transcriptional regulator involved in phenolic-acid metabolism.
- Sequence identity: 40.78%
- RMSD: 0.5 Å
- Predicted function: transcriptional regulator of phenolic acid
3.2.5. C1L0R8
Model built for residues 201–605 with 30.07% sequence identity and RMSD of 0.5 Å.
Conserved residues related to LTA synthesis and metal coordination support a role in lipoteichoic acid biosynthesis / sulfuric ester hydrolase-associated function.
- Sequence identity: 30.07%
- RMSD: 0.5 Å
3.2.6. C1L165
Model built for residues 5–176 with 41.28% sequence identity and RMSD of 0.2 Å.
Conserved residues support binding to poly-isoprenoid.
- Sequence identity: 41.28%
- RMSD: 0.2 Å
- Predicted function: poly-isoprenoid binding
3.2.7. C1L2E5
Model built for residues 1–158 with 22.50% sequence identity and RMSD of 3.919 Å.
Conserved metal-binding residues suggest possible phosphodiesterase activity, but low identity limits confidence.
- Sequence identity: 22.50%
- RMSD: 3.919 Å
- Confidence note: low-confidence functional inference
3.2.8. C1KZM7
Model built for residues 3–142 with 30.41% sequence identity and RMSD of 0.085 Å.
The study tentatively associates the protein with stress response and ectoine production involved in osmotic stress survival.
- Sequence identity: 30.41%
- RMSD: 0.085 Å
- Predicted function: possible stress protein / ectoine-related role
3.2.9. C1KYZ6
Model built for residues 42–196 with 12.35% sequence identity and RMSD of 5.687 Å.
Because of very low sequence identity and high RMSD, the authors regard this model as insufficiently reliable for function prediction.
- Sequence identity: 12.35%
- RMSD: 5.687 Å
- Confidence note: unreliable for functional inference
3.2.10. C1KYT3
Model built for residues 5–201 with 33.65% sequence identity and RMSD of 3.75 Å.
Conserved catalytic and substrate-binding residues suggest possible roles in differentiation, development, and regulation of cell proliferation.
- Sequence identity: 33.65%
- RMSD: 3.75 Å
3.2.11. C1KY51
Model built for residues 6–145 with 46.00% sequence identity and RMSD of 3.3 Å.
Conserved cysteines and Asp42 in the zinc-binding region support involvement in iron–sulfur cluster biology, electron transfer, or regulatory / sensing functions.
- Sequence identity: 46.00%
- RMSD: 3.3 Å
- Predicted function: iron ion / Fe-S cluster-related role
3.2.12. C1KYL6
Model built for residues 4–271 with 25.19% sequence identity and RMSD of 0.091 Å.
Conserved active-site residues support a HAD superfamily phosphatase function involved in phosphoryl-transfer and hydrolytic dephosphorylation.
- Sequence identity: 25.19%
- RMSD: 0.091 Å
- Predicted function: HAD superfamily phosphatase
3.2.13. C1KY50
Model built for residues 33–462 with 17.20% sequence identity and RMSD of 7.538 Å.
Conserved residues hint at an iron–sulfur cluster-related role, but the authors explicitly caution that the model is low-confidence and needs experimental validation.
- Sequence identity: 17.20%
- RMSD: 7.538 Å
- Confidence note: interpret with caution
5. Conclusion
The article concludes that sequence-based analysis allowed plausible biochemical function assignment for most of the 92 hypothetical proteins, while structural modeling enabled detailed structure-function interpretation for 13 selected proteins.
The work is positioned as a structural genomics contribution that improves understanding of Listeria monocytogenes pathogenesis and supports follow-up biochemical validation.
The authors also note that some proposed functions remain provisional and require further experimental work for precise confirmation.
- Hypothetical proteins analyzed: 92
- Proteins with detailed modeled structure-function analysis: 13
- Main limitation acknowledged: experimental validation still required
6. Statements and Declarations
The article includes closing statements for conflict of interest, data availability, funding, acknowledgements, supplementary materials, and author contributions.
- Conflict of Interest: There is no conflict of interest to declare.
- Data Availability Statement: The data supporting this study are provided in this article.
- Funding: None
- Acknowledgements: None
- Supplementary materials: None
Figures
The PDF includes nine named figures covering the computational workflow, mutation-count summaries, conservation mapping, structural models, and final network-style comparisons.
Figure 1
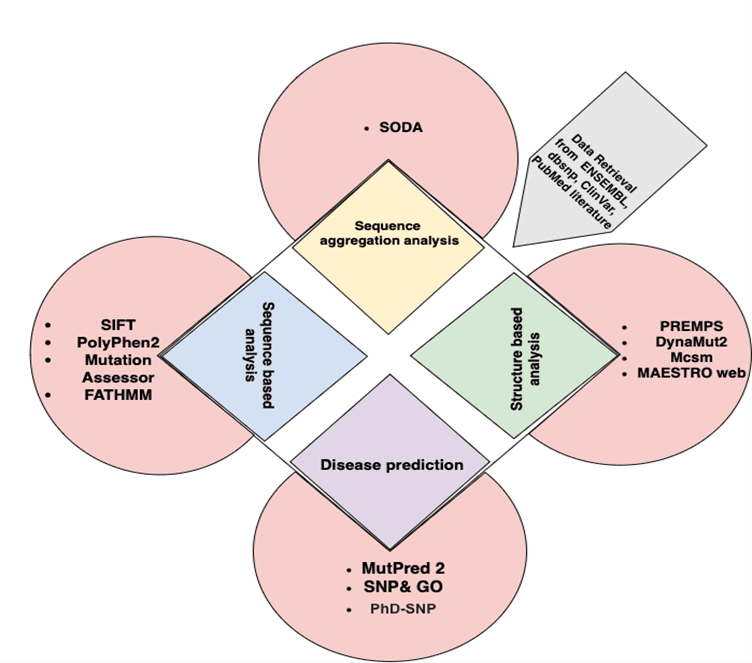

Figure 1. Outline of structure and function assignment to the hypothetical proteins.
Download figure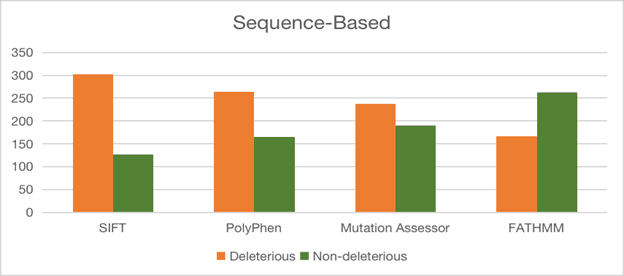
Figure 2. Structural superimposition of modeled proteins C1L1U8, C1KYM2, C1KYY1, C1L0M8, C1L0R8, and C1L165 with their respective template structures.
Download figure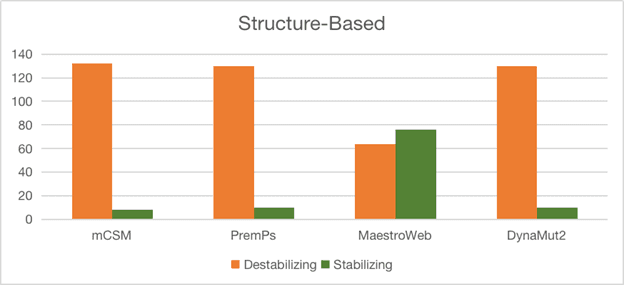
Figure 3. Structural superimposition of modeled proteins C1L2E5–C1KY50 with their respective template structures.
Download figureReferences
- 1
Chin, Ko-Hsin, Tsai, et al.
Reference cited in the paper.
2007 - 2
Fibriansah, G., Kovacs, A., Pool, T., et al.
PLoS ONE article cited in the paper.
PLoS ONE · 2012 - 3
Gury, J., Barthelmebs, L., Tran, N., Divies, C., & Cavin, J.
Applied and Environmental Microbiology article cited in the paper.
Appl Environ Microbiol · 2004 - 4
Handa, N., Terada, T., Doi-Katayama, Y., et al.
Protein Science article cited in the paper.
Protein Sci. · 2004 - 5
Liu, Jinyu, Oganesyan, et al.
Proteins article cited in the paper.
Proteins · 2005 - 6
Lu, Zhibing, Dunaway-Mariano, et al.
Proteins article cited in the paper.
Proteins · 2011 - 7
Rao, N.
Journal of Thoracic and Cardiovascular Surgery article cited in the paper.
J. Thorac. Cardiovasc. Surg. · 1989 - 8
Vorontsov, I., I., Minasov, et al.
Journal of Biological Chemistry article cited in the paper.
J. Biol. Chem. · 2011 - 9
Wada, K., Sumi, N., Nagai, R., et al.
Journal of Molecular Biology article cited in the paper.
J. Mol. Biol. · 2009 - 10
Xu, Yongbin, Sim, et al.
Biochemistry article cited in the paper.
Biochemistry · 2009


